diff --git a/1_WaPOR_download_colab/Download_WaPORv3_Data.ipynb b/1_WaPOR_download_colab/Download_WaPORv3_Data.ipynb
new file mode 100644
index 0000000..4fe82d1
--- /dev/null
+++ b/1_WaPOR_download_colab/Download_WaPORv3_Data.ipynb
@@ -0,0 +1,637 @@
+{
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "view-in-github",
+ "colab_type": "text"
+ },
+ "source": [
+ " "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "T9TpZLqEZ-Cc"
+ },
+ "source": [
+ "# Download WaPOR version 3 Data.\n",
+ "\n",
+ "[](https://colab.research.google.com/github/wateraccounting/WaPORMOOC/blob/main/1_WaPOR_download_colab/Download_WaPORv3_Data.ipynb?target=\"_blank\")\n",
+ "\n",
+ "To run the Notebook in Colab click [here](https://colab.research.google.com/github/wateraccounting/WaPORMOOC/blob/main/1_WaPOR_download_colab/Download_WaPORv3_Data.ipynb?target=\"_blank\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "VWN6GKMbZ-Ci"
+ },
+ "source": [
+ "---"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "GaiJb9WBsgBc"
+ },
+ "source": [
+ "\n",
+ "In this notebook we will use the [wapordl package](https://bitbucket.org/cioapps/wapordl/src/main/) made available by FAO to download WaPOR v3 data and download files to your local computer. This short [video](https://www.youtube.com/watch?v=ymBM_EVPEMQ) explains how to use this script. For details of how to use the downloaded WaPOR v3 data, refer to the [MOOC on Introduction to WaPORv3](https://ocw.un-ihe.org/course/view.php?id=263).\n",
+ "\n",
+ " \n",
+ "The steps are:\n",
+ "\n",
+ "0. Install and load necessary packages\n",
+ "1. Upload a geojson file to define the boundary of area of interest\n",
+ "2. Use the wapordl package to download raster files for each variable\n",
+ "3. Zip the rasters files and download them to local folder\n",
+ "\n",
+ "This script can be used without [mounting](https://colab.research.google.com/notebooks/snippets/drive.ipynb) Google colab with your Google Drive, if you want to use files stored in your Google Drive or save the files there, you have to mount the script first.\n",
+ "\n",
+ "**NOTE**: This script allows for downloading WaPOR data in separate tiffs using the WaPOR naming convention, removal of the scaling factor and creating data in the units per the temporal scale of the data (eg dekadal data is converted from mm/day to mm/dekade). For more information about WaPOR data formats check out this [video](https://www.youtube.com/watch?v=nm5IAy4MHg4). Removal of the scaling factor increases the file size, for more advanced users, check out the [wapordl package](https://bitbucket.org/cioapps/wapordl/src/main/) created by FAO for more ways to download WaPORv3 data (including an option for downloading 1 file with scaled values in multiple bands for saving space, do note to make sure the scaling factor is applied when utilising such data).\n",
+ "\n",
+ " © 2024 IHE Delft Licenced under CC BY SA Creative Commons"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "ypLr5xDlZ-Cj"
+ },
+ "source": [
+ "---"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "uirLoQBqA-Hs"
+ },
+ "source": [
+ "## **Step 0 - Install and load necessary packages**\n",
+ "As a check we also report the wapordl version you are using (to use to report an issue)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 1,
+ "metadata": {
+ "id": "3-CeI7rls2_4"
+ },
+ "outputs": [],
+ "source": [
+ "%%capture\n",
+ "!pip install --upgrade wapordl --quiet\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 2,
+ "metadata": {
+ "id": "74qweMAtU5C1",
+ "colab": {
+ "base_uri": "https://localhost:8080/"
+ },
+ "outputId": "58aa54cc-6034-436a-b327-5ed69dc9e331"
+ },
+ "outputs": [
+ {
+ "output_type": "stream",
+ "name": "stderr",
+ "text": [
+ "INFO: WaPORDL (`1.1.2`)\n"
+ ]
+ },
+ {
+ "output_type": "stream",
+ "name": "stdout",
+ "text": [
+ "1.1.2\n"
+ ]
+ }
+ ],
+ "source": [
+ "import wapordl\n",
+ "print(wapordl.__version__)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "5ZGHVNmRs6Bs"
+ },
+ "source": [
+ "#### Import and load required packages"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 3,
+ "metadata": {
+ "id": "T3EBB-5f7mcP"
+ },
+ "outputs": [],
+ "source": [
+ "# from osgeo import gdal\n",
+ "import glob\n",
+ "import os\n",
+ "from wapordl import wapor_map\n",
+ "from wapordl import wapor_ts"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "llH9UoaqIsAN"
+ },
+ "source": [
+ "\n",
+ "\n",
+ "---\n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "rtlfEcFHBTUy"
+ },
+ "source": [
+ "## **Step 1 - Upload a geojson file to define the boundary of area of interest**\n",
+ "Area of interest can be defined either by providing a geojson file or by bounding box (coordinates of the extent of the area of interest). In this step you can upload your geojson file. If you decide to use the bounding box approach you can skip this step.\n",
+ "\n",
+ "*NOTE*: wapordl only support geojson file format at the moment. If you have a file in other format, you can use QGIS to export it as geojson.\n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 7,
+ "metadata": {
+ "id": "DI1-Uc7JQETk",
+ "colab": {
+ "base_uri": "https://localhost:8080/",
+ "height": 73
+ },
+ "outputId": "1af10105-660b-42c3-8dd0-e4e8782d0f98"
+ },
+ "outputs": [
+ {
+ "output_type": "display_data",
+ "data": {
+ "text/plain": [
+ ""
+ ],
+ "text/html": [
+ "\n",
+ " \n",
+ " \n",
+ " "
+ ]
+ },
+ "metadata": {}
+ },
+ {
+ "output_type": "stream",
+ "name": "stdout",
+ "text": [
+ "Saving Bekaa.geojson to Bekaa.geojson\n"
+ ]
+ }
+ ],
+ "source": [
+ "#To upload file.\n",
+ "from google.colab import files\n",
+ "uploaded = files.upload()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "XPNh8UgrBt_A"
+ },
+ "source": [
+ "If the script doesnt run, please check if the geojson is in 2D format. (Shapely does currently not support 3D geometries, so the workaround is to [convert your 3D geometries to 2D](https://gis.stackexchange.com/questions/255540/convert-3d-shapefile-to-2d-in-qgis)). For more advanced users you can also link the script to a file stored in your google drive, but then you have to mount your script first."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "SAHKRORqItft"
+ },
+ "source": [
+ "\n",
+ "\n",
+ "---\n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "JTwt1Ng6CUaq"
+ },
+ "source": [
+ "## **Step 2 - Use the wapordl package to download raster files for each variable**\n",
+ "\n",
+ "To use the wapordl package to download WaPOR data, you need to 1) define the area of interest (either in geojson file or bounding box defined as [xmin, ymin, xmax, ymax]), 2) the variables to download and the time period, and 3) the folder where to save the downloaded files.\n",
+ "\n",
+ "The variables which can be downloaded include AETI (actual evapotranspiration and interception), T (transpiration), I (interception), E (evaporation), NPP (Net Primary Product), PCP (Precipitation), RET (Reference Evapotranspiration) and RSM (Relative Soil Moisture). There are 3 levels of WaPOR data (L1, L2 and L3) related to the resolution of the data and in terms of temporal resolution, Daily (E), Dekadal (D), Monthly (M) and Annual (A). Check in the [WaPOR portal](https://data.apps.fao.org/wapor/?lang=en) in which temporal resolution the data you want to download is available.\n",
+ "\n",
+ "Refer to the **[WaPOR overview](https://www.fao.org/in-action/remote-sensing-for-water-productivity/wapor-data/en)** to read more about the WaPOR data, its spatial and temporal coverage and the contents of the database. For the WaPOR variables naming convention, see the image below.\n",
+ "\n",
+ "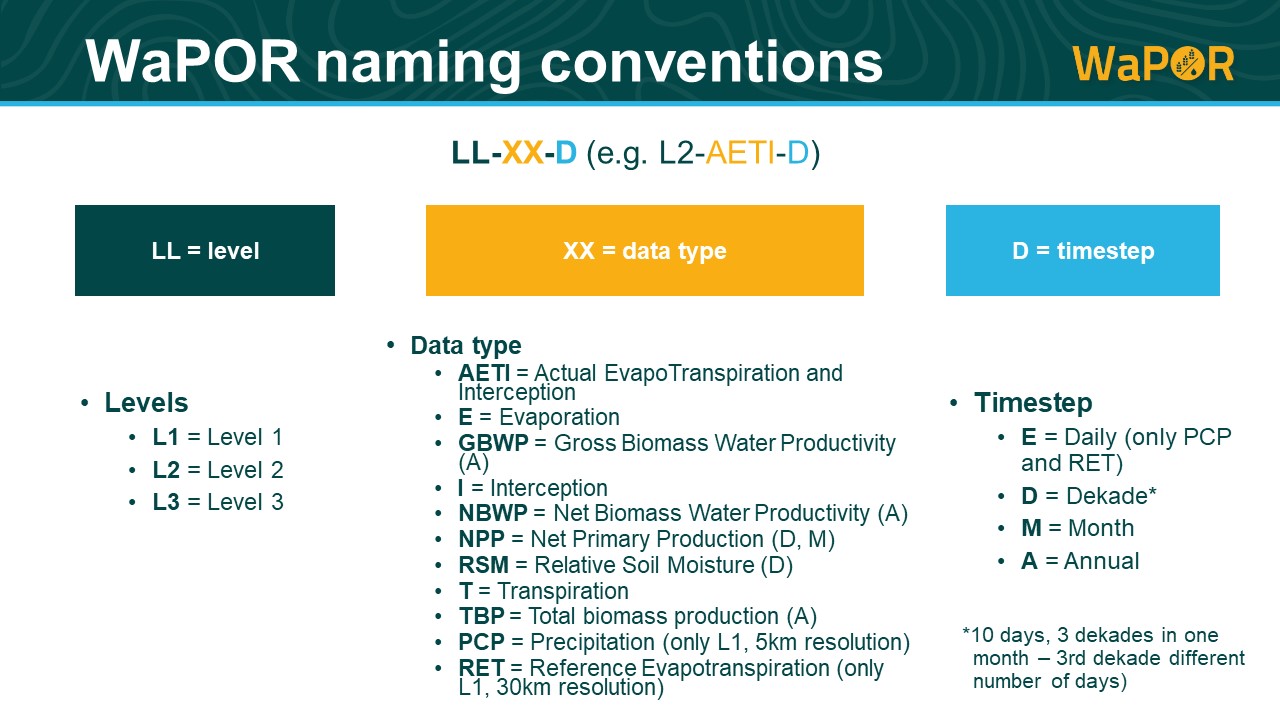"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 8,
+ "metadata": {
+ "id": "Fhrwhd5JQCLT",
+ "colab": {
+ "base_uri": "https://localhost:8080/"
+ },
+ "outputId": "cc3726fc-70fa-46a7-dea8-4f14aeeb5d2a"
+ },
+ "outputs": [
+ {
+ "output_type": "stream",
+ "name": "stderr",
+ "text": [
+ "INFO: Found 6 files for L2-AETI-A.\n",
+ "INFO: Units are already as requested, no conversion needed.\n",
+ "INFO: Splitting single GeoTIFF into multiple unscaled files.\n"
+ ]
+ }
+ ],
+ "source": [
+ "# 1) define region (remove comment (#) for the option you plan to use)\n",
+ "# define region using geojson file (if using a file stored in your google drive, add relative path here)\n",
+ "region = r\"/content/Bekaa.geojson\"\n",
+ "\n",
+ "# define region in bounding box\n",
+ "# region = [39.17, 8.32, 39.37, 8.50] # [xmin, ymin, xmax, ymax]\n",
+ "\n",
+ "# 2) define the variables and timeperiod to download\n",
+ "variables = [\"L2-AETI-A\"] #, \"L3-T-D\",\"L3-NPP-D\"]\n",
+ "period = [\"2018-01-01\", \"2023-12-31\"]\n",
+ "overview = \"NONE\"\n",
+ "\n",
+ "for var in variables:\n",
+ "\n",
+ " # 3) Define a folder to save the downloaded rasters (this will create a temporary folder\n",
+ " # called output, if you want the data to be stored somewhere else change the\n",
+ " # path, this requires mounting of the colab script)\n",
+ " folder = r\"/content/output\"+f\"/{var}\"\n",
+ " if not os.path.exists(folder):\n",
+ " os.makedirs(folder)\n",
+ "\n",
+ " if('-E' in var):\n",
+ " unit = \"day\"\n",
+ " elif('-D' in var):\n",
+ " unit = \"dekad\"\n",
+ " elif('-M' in var):\n",
+ " unit = \"month\"\n",
+ " elif ('-A' in var):\n",
+ " unit = \"year\"\n",
+ " else:\n",
+ " unit = \"none\"\n",
+ "\n",
+ " fps = wapor_map(region, var, period, folder, separate_unscale = True, unit_conversion = unit)\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "i2uG2PhMIvJ9"
+ },
+ "source": [
+ "The raster files can be found in the output folder, which can be manually downloaded. Alternatively go to step 3 to use a script for downloading. \n",
+ "\n",
+ "---\n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "jF9sxQquGtoH"
+ },
+ "source": [
+ "## **Step 3 - Zip the raster files and download them to local folder**\n",
+ "\n",
+ "This script will zip the contents of the output folder and download the zipped file to our local computer by running the following cell.\n",
+ "\n",
+ "*NOTE*: update the extensions if you have changed the output file location."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 9,
+ "metadata": {
+ "id": "9isKZdEm9mUs",
+ "colab": {
+ "base_uri": "https://localhost:8080/",
+ "height": 156
+ },
+ "outputId": "802125f6-1876-4435-e829-8a780b51e316"
+ },
+ "outputs": [
+ {
+ "output_type": "stream",
+ "name": "stdout",
+ "text": [
+ " adding: content/output/ (stored 0%)\n",
+ " adding: content/output/L2-AETI-A/ (stored 0%)\n",
+ " adding: content/output/L2-AETI-A/Bekaa_L2-AETI-A_NONE_year_2023-01-01.tif (deflated 12%)\n",
+ " adding: content/output/L2-AETI-A/Bekaa_L2-AETI-A_NONE_year_2018-01-01.tif (deflated 12%)\n",
+ " adding: content/output/L2-AETI-A/Bekaa_L2-AETI-A_NONE_year_2021-01-01.tif (deflated 12%)\n",
+ " adding: content/output/L2-AETI-A/Bekaa_L2-AETI-A_NONE_year_2022-01-01.tif (deflated 12%)\n",
+ " adding: content/output/L2-AETI-A/Bekaa_L2-AETI-A_NONE_year_2020-01-01.tif (deflated 12%)\n",
+ " adding: content/output/L2-AETI-A/Bekaa_L2-AETI-A_NONE_year_2019-01-01.tif (deflated 12%)\n"
+ ]
+ },
+ {
+ "output_type": "display_data",
+ "data": {
+ "text/plain": [
+ ""
+ ],
+ "application/javascript": [
+ "\n",
+ " async function download(id, filename, size) {\n",
+ " if (!google.colab.kernel.accessAllowed) {\n",
+ " return;\n",
+ " }\n",
+ " const div = document.createElement('div');\n",
+ " const label = document.createElement('label');\n",
+ " label.textContent = `Downloading \"${filename}\": `;\n",
+ " div.appendChild(label);\n",
+ " const progress = document.createElement('progress');\n",
+ " progress.max = size;\n",
+ " div.appendChild(progress);\n",
+ " document.body.appendChild(div);\n",
+ "\n",
+ " const buffers = [];\n",
+ " let downloaded = 0;\n",
+ "\n",
+ " const channel = await google.colab.kernel.comms.open(id);\n",
+ " // Send a message to notify the kernel that we're ready.\n",
+ " channel.send({})\n",
+ "\n",
+ " for await (const message of channel.messages) {\n",
+ " // Send a message to notify the kernel that we're ready.\n",
+ " channel.send({})\n",
+ " if (message.buffers) {\n",
+ " for (const buffer of message.buffers) {\n",
+ " buffers.push(buffer);\n",
+ " downloaded += buffer.byteLength;\n",
+ " progress.value = downloaded;\n",
+ " }\n",
+ " }\n",
+ " }\n",
+ " const blob = new Blob(buffers, {type: 'application/binary'});\n",
+ " const a = document.createElement('a');\n",
+ " a.href = window.URL.createObjectURL(blob);\n",
+ " a.download = filename;\n",
+ " div.appendChild(a);\n",
+ " a.click();\n",
+ " div.remove();\n",
+ " }\n",
+ " "
+ ]
+ },
+ "metadata": {}
+ },
+ {
+ "output_type": "display_data",
+ "data": {
+ "text/plain": [
+ ""
+ ],
+ "application/javascript": [
+ "download(\"download_86ff769e-864c-43c0-847d-daa4eb88b97e\", \"data.zip\", 2355846)"
+ ]
+ },
+ "metadata": {}
+ }
+ ],
+ "source": [
+ "!zip -r /content/data.zip /content/output\n",
+ "from google.colab import files\n",
+ "files.download(r'/content/data.zip')\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "id": "oEC1npPT7r_o"
+ },
+ "outputs": [],
+ "source": [
+ "# # if you want to delete a folder use the code below.\n",
+ "# !rm -rf /content/output\n",
+ "# !rm -rf /content/tifs\n",
+ "# !rm -rf /content/data.zip"
+ ]
+ }
+ ],
+ "metadata": {
+ "colab": {
+ "provenance": [],
+ "include_colab_link": true
+ },
+ "kernelspec": {
+ "display_name": "Python 3",
+ "name": "python3"
+ },
+ "language_info": {
+ "name": "python"
+ }
+ },
+ "nbformat": 4,
+ "nbformat_minor": 0
+}
\ No newline at end of file
diff --git a/Gemfile.lock b/Gemfile.lock
index e2e609e..f3e676d 100644
--- a/Gemfile.lock
+++ b/Gemfile.lock
@@ -6,8 +6,8 @@ GEM
minitest (~> 5.1)
thread_safe (~> 0.3, >= 0.3.4)
tzinfo (~> 1.1)
- addressable (2.7.0)
- public_suffix (>= 2.0.2, < 5.0)
+ addressable (2.8.1)
+ public_suffix (>= 2.0.2, < 6.0)
coffee-script (2.4.1)
coffee-script-source
execjs
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "T9TpZLqEZ-Cc"
+ },
+ "source": [
+ "# Download WaPOR version 3 Data.\n",
+ "\n",
+ "[](https://colab.research.google.com/github/wateraccounting/WaPORMOOC/blob/main/1_WaPOR_download_colab/Download_WaPORv3_Data.ipynb?target=\"_blank\")\n",
+ "\n",
+ "To run the Notebook in Colab click [here](https://colab.research.google.com/github/wateraccounting/WaPORMOOC/blob/main/1_WaPOR_download_colab/Download_WaPORv3_Data.ipynb?target=\"_blank\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "VWN6GKMbZ-Ci"
+ },
+ "source": [
+ "---"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "GaiJb9WBsgBc"
+ },
+ "source": [
+ "\n",
+ "In this notebook we will use the [wapordl package](https://bitbucket.org/cioapps/wapordl/src/main/) made available by FAO to download WaPOR v3 data and download files to your local computer. This short [video](https://www.youtube.com/watch?v=ymBM_EVPEMQ) explains how to use this script. For details of how to use the downloaded WaPOR v3 data, refer to the [MOOC on Introduction to WaPORv3](https://ocw.un-ihe.org/course/view.php?id=263).\n",
+ "\n",
+ " \n",
+ "The steps are:\n",
+ "\n",
+ "0. Install and load necessary packages\n",
+ "1. Upload a geojson file to define the boundary of area of interest\n",
+ "2. Use the wapordl package to download raster files for each variable\n",
+ "3. Zip the rasters files and download them to local folder\n",
+ "\n",
+ "This script can be used without [mounting](https://colab.research.google.com/notebooks/snippets/drive.ipynb) Google colab with your Google Drive, if you want to use files stored in your Google Drive or save the files there, you have to mount the script first.\n",
+ "\n",
+ "**NOTE**: This script allows for downloading WaPOR data in separate tiffs using the WaPOR naming convention, removal of the scaling factor and creating data in the units per the temporal scale of the data (eg dekadal data is converted from mm/day to mm/dekade). For more information about WaPOR data formats check out this [video](https://www.youtube.com/watch?v=nm5IAy4MHg4). Removal of the scaling factor increases the file size, for more advanced users, check out the [wapordl package](https://bitbucket.org/cioapps/wapordl/src/main/) created by FAO for more ways to download WaPORv3 data (including an option for downloading 1 file with scaled values in multiple bands for saving space, do note to make sure the scaling factor is applied when utilising such data).\n",
+ "\n",
+ " © 2024 IHE Delft Licenced under CC BY SA Creative Commons"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "ypLr5xDlZ-Cj"
+ },
+ "source": [
+ "---"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "uirLoQBqA-Hs"
+ },
+ "source": [
+ "## **Step 0 - Install and load necessary packages**\n",
+ "As a check we also report the wapordl version you are using (to use to report an issue)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 1,
+ "metadata": {
+ "id": "3-CeI7rls2_4"
+ },
+ "outputs": [],
+ "source": [
+ "%%capture\n",
+ "!pip install --upgrade wapordl --quiet\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 2,
+ "metadata": {
+ "id": "74qweMAtU5C1",
+ "colab": {
+ "base_uri": "https://localhost:8080/"
+ },
+ "outputId": "58aa54cc-6034-436a-b327-5ed69dc9e331"
+ },
+ "outputs": [
+ {
+ "output_type": "stream",
+ "name": "stderr",
+ "text": [
+ "INFO: WaPORDL (`1.1.2`)\n"
+ ]
+ },
+ {
+ "output_type": "stream",
+ "name": "stdout",
+ "text": [
+ "1.1.2\n"
+ ]
+ }
+ ],
+ "source": [
+ "import wapordl\n",
+ "print(wapordl.__version__)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "5ZGHVNmRs6Bs"
+ },
+ "source": [
+ "#### Import and load required packages"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 3,
+ "metadata": {
+ "id": "T3EBB-5f7mcP"
+ },
+ "outputs": [],
+ "source": [
+ "# from osgeo import gdal\n",
+ "import glob\n",
+ "import os\n",
+ "from wapordl import wapor_map\n",
+ "from wapordl import wapor_ts"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "llH9UoaqIsAN"
+ },
+ "source": [
+ "\n",
+ "\n",
+ "---\n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "rtlfEcFHBTUy"
+ },
+ "source": [
+ "## **Step 1 - Upload a geojson file to define the boundary of area of interest**\n",
+ "Area of interest can be defined either by providing a geojson file or by bounding box (coordinates of the extent of the area of interest). In this step you can upload your geojson file. If you decide to use the bounding box approach you can skip this step.\n",
+ "\n",
+ "*NOTE*: wapordl only support geojson file format at the moment. If you have a file in other format, you can use QGIS to export it as geojson.\n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 7,
+ "metadata": {
+ "id": "DI1-Uc7JQETk",
+ "colab": {
+ "base_uri": "https://localhost:8080/",
+ "height": 73
+ },
+ "outputId": "1af10105-660b-42c3-8dd0-e4e8782d0f98"
+ },
+ "outputs": [
+ {
+ "output_type": "display_data",
+ "data": {
+ "text/plain": [
+ ""
+ ],
+ "text/html": [
+ "\n",
+ " \n",
+ " \n",
+ " "
+ ]
+ },
+ "metadata": {}
+ },
+ {
+ "output_type": "stream",
+ "name": "stdout",
+ "text": [
+ "Saving Bekaa.geojson to Bekaa.geojson\n"
+ ]
+ }
+ ],
+ "source": [
+ "#To upload file.\n",
+ "from google.colab import files\n",
+ "uploaded = files.upload()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "XPNh8UgrBt_A"
+ },
+ "source": [
+ "If the script doesnt run, please check if the geojson is in 2D format. (Shapely does currently not support 3D geometries, so the workaround is to [convert your 3D geometries to 2D](https://gis.stackexchange.com/questions/255540/convert-3d-shapefile-to-2d-in-qgis)). For more advanced users you can also link the script to a file stored in your google drive, but then you have to mount your script first."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "SAHKRORqItft"
+ },
+ "source": [
+ "\n",
+ "\n",
+ "---\n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "JTwt1Ng6CUaq"
+ },
+ "source": [
+ "## **Step 2 - Use the wapordl package to download raster files for each variable**\n",
+ "\n",
+ "To use the wapordl package to download WaPOR data, you need to 1) define the area of interest (either in geojson file or bounding box defined as [xmin, ymin, xmax, ymax]), 2) the variables to download and the time period, and 3) the folder where to save the downloaded files.\n",
+ "\n",
+ "The variables which can be downloaded include AETI (actual evapotranspiration and interception), T (transpiration), I (interception), E (evaporation), NPP (Net Primary Product), PCP (Precipitation), RET (Reference Evapotranspiration) and RSM (Relative Soil Moisture). There are 3 levels of WaPOR data (L1, L2 and L3) related to the resolution of the data and in terms of temporal resolution, Daily (E), Dekadal (D), Monthly (M) and Annual (A). Check in the [WaPOR portal](https://data.apps.fao.org/wapor/?lang=en) in which temporal resolution the data you want to download is available.\n",
+ "\n",
+ "Refer to the **[WaPOR overview](https://www.fao.org/in-action/remote-sensing-for-water-productivity/wapor-data/en)** to read more about the WaPOR data, its spatial and temporal coverage and the contents of the database. For the WaPOR variables naming convention, see the image below.\n",
+ "\n",
+ ""
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 8,
+ "metadata": {
+ "id": "Fhrwhd5JQCLT",
+ "colab": {
+ "base_uri": "https://localhost:8080/"
+ },
+ "outputId": "cc3726fc-70fa-46a7-dea8-4f14aeeb5d2a"
+ },
+ "outputs": [
+ {
+ "output_type": "stream",
+ "name": "stderr",
+ "text": [
+ "INFO: Found 6 files for L2-AETI-A.\n",
+ "INFO: Units are already as requested, no conversion needed.\n",
+ "INFO: Splitting single GeoTIFF into multiple unscaled files.\n"
+ ]
+ }
+ ],
+ "source": [
+ "# 1) define region (remove comment (#) for the option you plan to use)\n",
+ "# define region using geojson file (if using a file stored in your google drive, add relative path here)\n",
+ "region = r\"/content/Bekaa.geojson\"\n",
+ "\n",
+ "# define region in bounding box\n",
+ "# region = [39.17, 8.32, 39.37, 8.50] # [xmin, ymin, xmax, ymax]\n",
+ "\n",
+ "# 2) define the variables and timeperiod to download\n",
+ "variables = [\"L2-AETI-A\"] #, \"L3-T-D\",\"L3-NPP-D\"]\n",
+ "period = [\"2018-01-01\", \"2023-12-31\"]\n",
+ "overview = \"NONE\"\n",
+ "\n",
+ "for var in variables:\n",
+ "\n",
+ " # 3) Define a folder to save the downloaded rasters (this will create a temporary folder\n",
+ " # called output, if you want the data to be stored somewhere else change the\n",
+ " # path, this requires mounting of the colab script)\n",
+ " folder = r\"/content/output\"+f\"/{var}\"\n",
+ " if not os.path.exists(folder):\n",
+ " os.makedirs(folder)\n",
+ "\n",
+ " if('-E' in var):\n",
+ " unit = \"day\"\n",
+ " elif('-D' in var):\n",
+ " unit = \"dekad\"\n",
+ " elif('-M' in var):\n",
+ " unit = \"month\"\n",
+ " elif ('-A' in var):\n",
+ " unit = \"year\"\n",
+ " else:\n",
+ " unit = \"none\"\n",
+ "\n",
+ " fps = wapor_map(region, var, period, folder, separate_unscale = True, unit_conversion = unit)\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "i2uG2PhMIvJ9"
+ },
+ "source": [
+ "The raster files can be found in the output folder, which can be manually downloaded. Alternatively go to step 3 to use a script for downloading. \n",
+ "\n",
+ "---\n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "id": "jF9sxQquGtoH"
+ },
+ "source": [
+ "## **Step 3 - Zip the raster files and download them to local folder**\n",
+ "\n",
+ "This script will zip the contents of the output folder and download the zipped file to our local computer by running the following cell.\n",
+ "\n",
+ "*NOTE*: update the extensions if you have changed the output file location."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 9,
+ "metadata": {
+ "id": "9isKZdEm9mUs",
+ "colab": {
+ "base_uri": "https://localhost:8080/",
+ "height": 156
+ },
+ "outputId": "802125f6-1876-4435-e829-8a780b51e316"
+ },
+ "outputs": [
+ {
+ "output_type": "stream",
+ "name": "stdout",
+ "text": [
+ " adding: content/output/ (stored 0%)\n",
+ " adding: content/output/L2-AETI-A/ (stored 0%)\n",
+ " adding: content/output/L2-AETI-A/Bekaa_L2-AETI-A_NONE_year_2023-01-01.tif (deflated 12%)\n",
+ " adding: content/output/L2-AETI-A/Bekaa_L2-AETI-A_NONE_year_2018-01-01.tif (deflated 12%)\n",
+ " adding: content/output/L2-AETI-A/Bekaa_L2-AETI-A_NONE_year_2021-01-01.tif (deflated 12%)\n",
+ " adding: content/output/L2-AETI-A/Bekaa_L2-AETI-A_NONE_year_2022-01-01.tif (deflated 12%)\n",
+ " adding: content/output/L2-AETI-A/Bekaa_L2-AETI-A_NONE_year_2020-01-01.tif (deflated 12%)\n",
+ " adding: content/output/L2-AETI-A/Bekaa_L2-AETI-A_NONE_year_2019-01-01.tif (deflated 12%)\n"
+ ]
+ },
+ {
+ "output_type": "display_data",
+ "data": {
+ "text/plain": [
+ ""
+ ],
+ "application/javascript": [
+ "\n",
+ " async function download(id, filename, size) {\n",
+ " if (!google.colab.kernel.accessAllowed) {\n",
+ " return;\n",
+ " }\n",
+ " const div = document.createElement('div');\n",
+ " const label = document.createElement('label');\n",
+ " label.textContent = `Downloading \"${filename}\": `;\n",
+ " div.appendChild(label);\n",
+ " const progress = document.createElement('progress');\n",
+ " progress.max = size;\n",
+ " div.appendChild(progress);\n",
+ " document.body.appendChild(div);\n",
+ "\n",
+ " const buffers = [];\n",
+ " let downloaded = 0;\n",
+ "\n",
+ " const channel = await google.colab.kernel.comms.open(id);\n",
+ " // Send a message to notify the kernel that we're ready.\n",
+ " channel.send({})\n",
+ "\n",
+ " for await (const message of channel.messages) {\n",
+ " // Send a message to notify the kernel that we're ready.\n",
+ " channel.send({})\n",
+ " if (message.buffers) {\n",
+ " for (const buffer of message.buffers) {\n",
+ " buffers.push(buffer);\n",
+ " downloaded += buffer.byteLength;\n",
+ " progress.value = downloaded;\n",
+ " }\n",
+ " }\n",
+ " }\n",
+ " const blob = new Blob(buffers, {type: 'application/binary'});\n",
+ " const a = document.createElement('a');\n",
+ " a.href = window.URL.createObjectURL(blob);\n",
+ " a.download = filename;\n",
+ " div.appendChild(a);\n",
+ " a.click();\n",
+ " div.remove();\n",
+ " }\n",
+ " "
+ ]
+ },
+ "metadata": {}
+ },
+ {
+ "output_type": "display_data",
+ "data": {
+ "text/plain": [
+ ""
+ ],
+ "application/javascript": [
+ "download(\"download_86ff769e-864c-43c0-847d-daa4eb88b97e\", \"data.zip\", 2355846)"
+ ]
+ },
+ "metadata": {}
+ }
+ ],
+ "source": [
+ "!zip -r /content/data.zip /content/output\n",
+ "from google.colab import files\n",
+ "files.download(r'/content/data.zip')\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "id": "oEC1npPT7r_o"
+ },
+ "outputs": [],
+ "source": [
+ "# # if you want to delete a folder use the code below.\n",
+ "# !rm -rf /content/output\n",
+ "# !rm -rf /content/tifs\n",
+ "# !rm -rf /content/data.zip"
+ ]
+ }
+ ],
+ "metadata": {
+ "colab": {
+ "provenance": [],
+ "include_colab_link": true
+ },
+ "kernelspec": {
+ "display_name": "Python 3",
+ "name": "python3"
+ },
+ "language_info": {
+ "name": "python"
+ }
+ },
+ "nbformat": 4,
+ "nbformat_minor": 0
+}
\ No newline at end of file
diff --git a/Gemfile.lock b/Gemfile.lock
index e2e609e..f3e676d 100644
--- a/Gemfile.lock
+++ b/Gemfile.lock
@@ -6,8 +6,8 @@ GEM
minitest (~> 5.1)
thread_safe (~> 0.3, >= 0.3.4)
tzinfo (~> 1.1)
- addressable (2.7.0)
- public_suffix (>= 2.0.2, < 5.0)
+ addressable (2.8.1)
+ public_suffix (>= 2.0.2, < 6.0)
coffee-script (2.4.1)
coffee-script-source
execjs