-
Notifications
You must be signed in to change notification settings - Fork 0
Expand file tree
/
Copy path01-Fundamentals.Rmd
More file actions
325 lines (214 loc) · 10.3 KB
/
01-Fundamentals.Rmd
File metadata and controls
325 lines (214 loc) · 10.3 KB
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
110
111
112
113
114
115
116
117
118
119
120
121
122
123
124
125
126
127
128
129
130
131
132
133
134
135
136
137
138
139
140
141
142
143
144
145
146
147
148
149
150
151
152
153
154
155
156
157
158
159
160
161
162
163
164
165
166
167
168
169
170
171
172
173
174
175
176
177
178
179
180
181
182
183
184
185
186
187
188
189
190
191
192
193
194
195
196
197
198
199
200
201
202
203
204
205
206
207
208
209
210
211
212
213
214
215
216
217
218
219
220
221
222
223
224
225
226
227
228
229
230
231
232
233
234
235
236
237
238
239
240
241
242
243
244
245
246
247
248
249
250
251
252
253
254
255
256
257
258
259
260
261
262
263
264
265
266
267
268
269
270
271
272
273
274
275
276
277
278
279
280
281
282
283
284
285
286
287
288
289
290
291
292
293
294
295
296
297
298
299
300
301
302
303
304
305
306
307
308
309
310
311
312
313
314
315
316
317
318
319
320
321
322
323
324
325
# Fundamentals
```{r, echo=F, message=F, warning=F}
install.packages("palmerpenguins")
library(palmerpenguins)
library(tidyverse)
```
Welcome to Intermediate R! Each week, we cover a chapter, which consists of a lesson and exercise. In the first week, we go over the goals of the course, and review data structures and data types that you have seen before from [Intro to R](https://hutchdatascience.org/Intro_to_R/). We also look at some new data structures and more properties of data structures.
In [Intro to R](https://hutchdatascience.org/Intro_to_R/), you learned how to do basic data analysis such as subsetting a dataframe, looking at summary statistics, and visualizing your data. This was done in the context of a clean, Tidy dataframe. In this course, we focus on working with data "from the wild", in which the data comes in a more messy, un-Tidy form. Let's see what we will learn in the next 6 weeks together:
## Goals of this course
- Continue building *programming fundamentals*: How to use complex data structures, use and create custom functions, and how to iterate repeated tasks.
- Continue exploration of *data science fundamentals*: how to clean messy data to a Tidy form for analysis.
- At the end of the course, you will be able to: conduct a full analysis in the data science workflow (minus model).
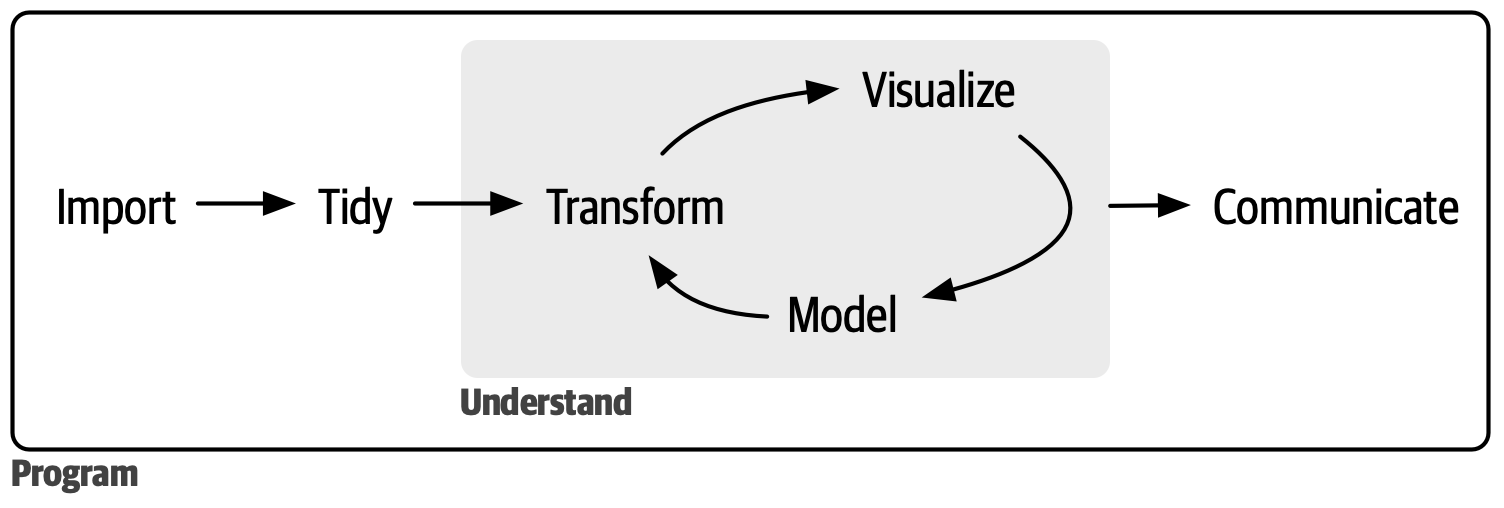{width="450"}
To get started, let's recall the fundamental data types in R:
## Data types in R
- Numeric: 18, -21, 65, 1.25
- Character: "ATCG", "Whatever", "948-293-0000"
- Logical: TRUE, FALSE
- Missing values: `NA`
And the fundamental data structures: in this course, we will learn more about a new, flexible data structure called a **List**. We also lightly introduce *Factor* and *Matrix*, but they will not be used for the rest of the course.
## Data structures
- Vector
- *Factor*
- Dataframe
- **List**
- *Matrix*
## Vector
We know what an **(atomic) vector** is: it can contains a data type, and all elements must be the same data type. If a vector consists of only numeric data, then it is a Numeric Vector, etc. We organize vector subtypes by the following graphic:
.](https://d33wubrfki0l68.cloudfront.net/eb6730b841e32292d9ff36b33a590e24b6221f43/57192/diagrams/vectors/summary-tree-atomic.png){width="400"}
Within the Numeric type that we are familiar with, there are more specific types: *Integer* consists of whole number values, and *Double* consists of decimal values. Most of the time we only need to consider Numeric types, but once in a while we need to be more specific.
Now that we have distinguished vector subtypes, it is important to examine what a vector is by inspection:
- We can test whether a vector is a certain type with `is.___()` functions, such as `is.character()`.
```{r}
is.character(c("hello", "there"))
is.character(c(1, 3, 5, 7))
```
- We can also test for missing data `NA` for any types of vector: The test will return a vector testing each element, because `NA` can be mixed into other values:
```{r}
is.na(c(34, NA))
```
We can **coerce** vectors from one type to the other with `as.___()` functions, such as `as.numeric()`.
```{r}
as.numeric(c("23", "45"))
as.numeric(c(TRUE, FALSE))
```
This is very common in data cleaning, when we load in data and they assigned to the wrong data type.
Sometimes, a data structure may have metadata **attributes** associated with them. This gives us more information about the data structure, but doesn't contain the important data.
For instance, a common attribute is **names,** which can attached to vectors.
```{r}
x = c(1, 2, 3)
names(x) = c("a", "b", "c")
x
```
](https://d33wubrfki0l68.cloudfront.net/1140c34226b3b04438aec65c8fc6b28758d8c091/1748a/diagrams/vectors/attr-names-2.png)
We can look for more general attributes beyond names via the `attributes()` function:
```{r}
attributes(x)
```
Now, let's review the ways one can subset a vector. Here are three ways:
1. Positive numeric vector
```{r}
data = c(2, 4, -1, -3, 2, -1, 10)
data[c(1, 3, 5)]
```
2. Negative numeric vector performs *exclusion*
```{r}
data[c(-1, -2)]
```
3. Logical vector
```{r}
data[c(TRUE, FALSE, TRUE, FALSE, TRUE, FALSE, FALSE)]
```
In practice, we often subset a vector implicitly, via some kind of criteria. Here is a [review of implicit subsetting from Intro to R](https://hutchdatascience.org/Intro_to_R/working-with-data-structures.html#subsetting-vectors-implicitly). Let's review implicit vector subsetting below:
1. How do you subset the following vector so that it only has positive values?
```{r}
data = c(2, 4, -1, -3, 2, -1, 10)
```
```{r}
data[data > 0]
```
2. How do you subset the following vector so that it has doesn't have the character "temp"?
```{r}
chars = c("temp", "object", "temp", "wish", "bumblebee", "temp")
```
```{r}
chars[chars != "temp"]
```
3. How do you subset the following vector so that it has no `NA` values?
```{r}
vec_with_NA = c(2, 4, NA, NA, 3, NA)
```
```{r}
vec_with_NA[!is.na(vec_with_NA)]
```
## Factors
Factors are a type of vector that holds categorical information, such as sex, gender, or cancer subtype. They are useful for:
- When you know you have a fixed number of categories.
- When you want to display character vectors in a non-alphabetical order, which is common in plotting.
- Inputs for statistical models, as factors are a special type of numerical vectors.
Let's take a look at Factors in practice:
```{r}
place = factor(c("first", "third", "third", "second", "second", "fourth"))
place
```
```{r}
df = data.frame(p = place)
ggplot(df) + geom_bar(aes(x = p))
```
We can construct Ordered Factors:
```{r}
place = ordered(c("first", "third", "third", "second","second", "fourth"), levels = c("first", "second", "third", "fourth"))
place
df = data.frame(p = place)
ggplot(df) + geom_bar(aes(x = p))
```
## Dataframes
Usually, we load in a Dataframe from a spreadsheet or a package, but we can create a new dataframe by using vectors of the same length via the `data.frame()` function:
```{r}
df = data.frame(x = 1:3, y = c("cup", "mug", "jar"))
df
```
We have attributes for Dataframes. The most important attribute is names, which correspond to the column names of a Dataframe. You have been using it for a while already!
```{r}
attributes(df)
```
We can directly access the names attribute via `names()` or `colnames()`:
```{r}
names(df)
```
Here is another example:
```{r}
library(palmerpenguins)
attributes(penguins)
```
Some notes about the other attributes:
- Sometimes, Dataframes will be in a format called "[tibble](https://tibble.tidyverse.org/)", as shown in the `penguins` class names as "tbl_df", and "tbl".
- Row names are not commonly used. Here is a [reason](https://adv-r.hadley.nz/vectors-chap.html#rownames).
Let's review how to subset Dataframes. There are many ways to do it, and here are just some opinionated ways of doing it for this class.
*Getting one single column:*
```{r}
penguins$bill_length_mm
```
*I want to select columns `bill_length_mm`, `bill_depth_mm`, `species`, and filter for `species` that are "Gentoo":*
```{r}
penguins_select = select(penguins, bill_length_mm, bill_depth_mm, species)
penguins_gentoo = filter(penguins_select, species == "Gentoo")
```
or
```{r}
penguins_select_2 = penguins[, c("bill_length_mm", "bill_depth_mm", "species")]
penguins_gentoo_2 = penguins_select_2[penguins$species == "Gentoo" ,]
```
or
```{r}
penguins_gentoo_2 = penguins_select_2[penguins$species == "Gentoo", c("bill_length_mm", "bill_depth_mm", "species")]
```
*I want to filter out rows that have `NA`s in the column `bill_length_mm`:*
```{r}
penguins_clean = filter(penguins, !is.na(bill_length_mm))
```
or
```{r}
penguins_clean = penguins[!is.na(penguins$bill_depth_mm) ,]
```
## Lists
Lists are the most flexible data structure in R, as they can contain a flexible amount and type of information. They operate similarly as vectors as they group data into one dimension, but each element of a list can be any data type *or data structure*!
```{r}
l1 = list(
1:3,
"a",
c(TRUE, FALSE, TRUE),
c(2.3, 5.9)
)
```
](https://d33wubrfki0l68.cloudfront.net/9628eed602df6fd55d9bced4fba0a5a85d93db8a/36c16/diagrams/vectors/list.png)
Unlike vectors, you access the elements of a list via the double bracket `[[]]`. You access a smaller list with single bracket `[]`. (More discussion on the different uses of the bracket [here](https://adv-r.hadley.nz/subsetting.html#subset-single) and [here](https://stackoverflow.com/questions/1169456/the-difference-between-bracket-and-double-bracket-for-accessing-the-el).)
Use `unlist()` to coerce a list into a vector. Notice all the automatic coersion that happened for the elements.
```{r}
unlist(l1)
```
We can give the attribute **names** to lists:
```{r}
l1 = list(
ranking = 1:3,
name = "a",
success = c(TRUE, FALSE, TRUE),
score = c(2.3, 5.9)
)
#or
names(l1) = c("ranking", "name", "success", "score")
```
And access named elements of lists via the `$` operation:
```{r}
l1$score
```
Therefore, `l1$score` is the same as `l1[[4]]` and is the same as `l1[["score"]]`.
Here's an interesting connection between Lists and Dataframes that we will make use of later on in this course: A Dataframe is just a named list of vectors of same length with additional **attributes** of (column) `names` and `row.names`!
## Matrix
A matrix holds information of the same data type in two dimensions - it's like a two dimensional vector. Matricies are most often used in statistical computing and matrix algebra, such as creating a design matrix. They are often created by taking a vector and reshaping it with a set number of rows and columns, or converting from a dataframe with only one data type.
```{r}
my_matrix = matrix(1:10, nrow = 2)
my_matrix
```
You access elements of a matrix similar to that of a dataframe's indexing:
```{r}
#column 3
my_matrix[, 3]
#row 2
my_matrix[2 ,]
#column 3, row 2
my_matrix[2, 3]
```
## Exercises
You can find [exercises and solutions on Posit Cloud](https://posit.cloud/content/8236252), or on [GitHub](https://github.com/fhdsl/Intermediate_R_Exercises).