|
1 | 1 | # Windowed Multipole Library Format |
2 | 2 |
|
3 | | -**/version** (*char[]*) |
| 3 | +The current version of the statepoint file format is 1.1. |
4 | 4 |
|
5 | | - The format version of the file. The current version is "v1.0" |
| 5 | ++ **/** |
6 | 6 |
|
7 | | -**/nuclide/** |
| 7 | + - *Attributes* : |
8 | 8 |
|
9 | | -- **broaden_poly** (*int[]*) |
| 9 | + - **filetype** (*char[]*) |
10 | 10 |
|
11 | | - If 1, Doppler broaden curve fit for window with corresponding index. |
12 | | - If 0, do not. |
| 11 | + String indicating the type of file. |
13 | 12 |
|
14 | | -- **curvefit** (*double[][][]*) |
| 13 | + - **version** (*int[2]*) |
15 | 14 |
|
16 | | - Curve fit coefficients. Indexed by (reaction type, coefficient index, |
17 | | - window index). |
| 15 | + Major and minor version of the data. |
18 | 16 |
|
19 | | -- **data** (*complex[][]*) |
| 17 | ++ **/\<nuclide name\>/** |
20 | 18 |
|
21 | | - Complex poles and residues. Each pole has a corresponding set of |
22 | | - residues. For example, the i-th pole and corresponding residues |
23 | | - are stored as |
| 19 | + - *Datasets* : |
24 | 20 |
|
25 | | -  |
| 21 | + - **broaden_poly** (*int[]*) |
26 | 22 |
|
27 | | - The residues are in the order: scattering, absorption, fission. Complex |
28 | | - numbers are stored by forming a type with "r" and "i" identifiers, |
29 | | - similar to how [h5py] does it. |
| 23 | + If 1, Doppler broaden curve fit for window with corresponding index. |
| 24 | + If 0, do not. |
30 | 25 |
|
31 | | -- **E_max** (*double*) |
| 26 | + - **curvefit** (*double[][][]*) |
32 | 27 |
|
33 | | - Highest energy the windowed multipole part of the library is valid for. |
| 28 | + Curve fit coefficients. Indexed by (reaction type, coefficient index, |
| 29 | + window index). |
34 | 30 |
|
35 | | -- **E_min** (*double*) |
| 31 | + - **data** (*complex[][]*) |
36 | 32 |
|
37 | | - Lowest energy the windowed multipole part of the library is valid for. |
| 33 | + Complex poles and residues. Each pole has a corresponding set of |
| 34 | + residues. For example, the i-th pole and corresponding residues |
| 35 | + are stored as |
38 | 36 |
|
39 | | -- **spacing** (*double*) |
| 37 | +  |
40 | 38 |
|
41 | | -  |
| 39 | + The residues are in the order: scattering, absorption, fission. Complex |
| 40 | + numbers are stored by forming a type with "r" and "i" identifiers, |
| 41 | + similar to how [h5py] does it. |
42 | 42 |
|
43 | | - Where 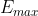 is the |
44 | | - maximum energy the windows go up to. |
45 | | -  is the minimum |
46 | | - energy and equivalent to ``E_min``, and 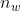 |
47 | | - is the number of windows, given by ``windows``. |
| 43 | + - **E_max** (*double*) |
48 | 44 |
|
49 | | -- **sqrtAWR** (*double*) |
| 45 | + Highest energy the windowed multipole part of the library is valid for. |
50 | 46 |
|
51 | | - Square root of the atomic weight ratio. |
| 47 | + - **E_min** (*double*) |
52 | 48 |
|
53 | | -- **windows** (*int[][]*) |
| 49 | + Lowest energy the windowed multipole part of the library is valid for. |
54 | 50 |
|
55 | | - The poles to start from and end at for each window. windows[i, 0] and |
56 | | - windows[i, 1] are, respectively, the indexes (1-based) of the first and last |
57 | | - pole in window i. |
| 51 | + - **spacing** (*double*) |
| 52 | + |
| 53 | +  |
| 54 | + |
| 55 | + Where 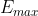 is the |
| 56 | + maximum energy the windows go up to. |
| 57 | +  is the minimum |
| 58 | + energy and equivalent to ``E_min``, and 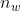 |
| 59 | + is the number of windows, given by ``windows``. |
| 60 | + |
| 61 | + - **sqrtAWR** (*double*) |
| 62 | + |
| 63 | + Square root of the atomic weight ratio. |
| 64 | + |
| 65 | + - **windows** (*int[][]*) |
| 66 | + |
| 67 | + The poles to start from and end at for each window. windows[i, 0] and |
| 68 | + windows[i, 1] are, respectively, the indexes (1-based) of the first and last |
| 69 | + pole in window i. |
58 | 70 |
|
59 | 71 | [h5py]: http://docs.h5py.org/en/latest/ |
0 commit comments