Releases: ArcInstitute/evo2
evo2 v0.5.0: Evo 2 20B release
Evo 2 20B: strong performance with half the GPU footprint
Garyk Brixi, Daniel Chang, Brian Hie
We are releasing Evo 2 20B, a model which approaches the performance of Evo 2 40B with half of the parameters, enabling it to run on a single H100 GPU.
Evo 2 40B was our strongest model but required over 80 GB of GPU memory. While the Nvidia NIM enables cloud inference of Evo 2, many applications are easier with a local model. Evo 2 20B makes this possible.
Evo 2 20B is a drop in alternative for other Evo 2 models and can be run using the Evo 2 library and fine-tuned using BioNemo or Savanna.
Quickstart
from evo2 import Evo2
model = Evo2('evo2_20b')
output = model.generate(prompt_seqs=["ACGT"], n_tokens=100, temperature=0.7, top_k=4)
print(output.sequences[0])
Evaluations
We evaluate Evo 2 20B on variant effect prediction and generation tasks. For variant effect prediction, we perform zero-shot evaluation of TraitGym Mendelian and Complex Traits, as well as BRCA2 SGE, and BRCA1 DMS. For generative tasks we perform gene completion. Methods for BRCA1, BRCA2, and gene completion evals are described in the Evo 2 preprint.
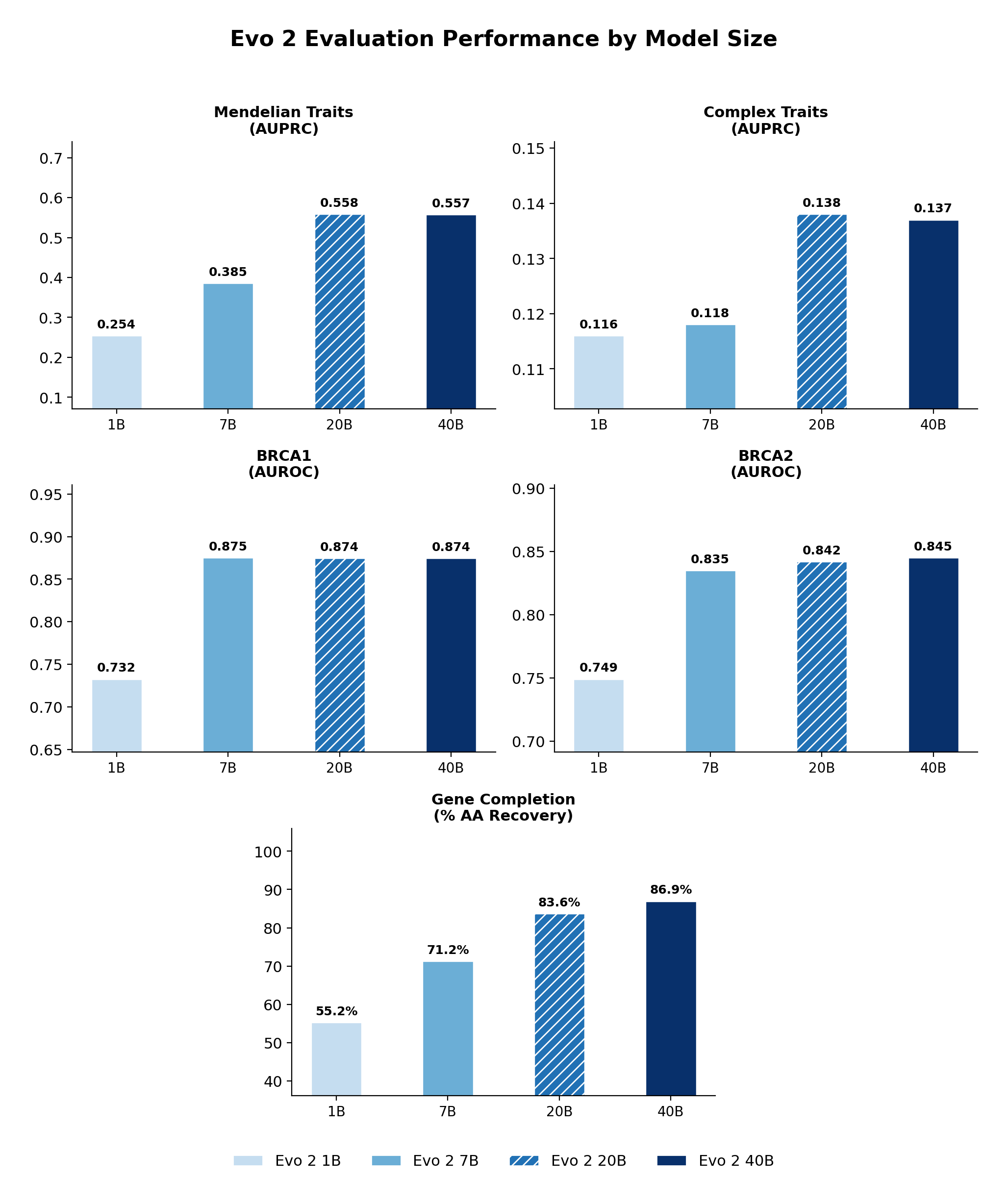
Across tasks, Evo 2 20B performs comparably to Evo 2 40B.
Evo 2 model comparison across benchmarks. Performance of Evo 2 1B, 7B, 20B, and 40B on variant effect prediction (TraitGym, BRCA1, BRCA2) and gene completion. Evo 2 20B closely matches Evo 2 40B across all tasks.
See here for more detailed baselines.
Hardware & Performance
Evo 2 20B weights are 38GB on GPU, compared to 80 for Evo 2 40B. Note that like Evo 2 40B, Evo 2 20B does need an Nvidia Hopper GPU for correctness due to requiring Transformer Engine's FP8 linear layers.
Benchmarked on NVIDIA H100 80GB HBM3:
| Evo 2 20B | Evo 2 40B | |
|---|---|---|
| GPUs required | 1× H100 | 2× H100 |
| Weight memory | 38 GB | 80 GB |
| Peak memory (8kb inference) | 46 GB | 95 GB |
| Score 8kb sequence | 0.7s | 1.5s |
| Generate 1kb | 20s | 42s |
Methods
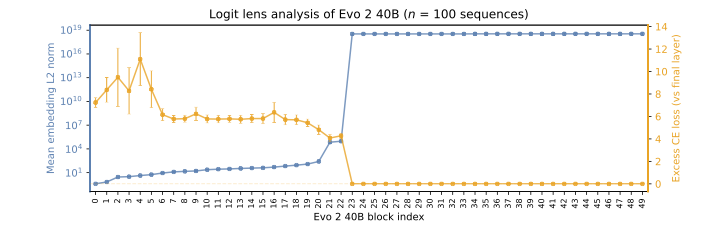
Evo 2 20B was created through model surgery of Evo 2 40B. Using logit lens analysis, we identified layers that could be removed without significantly affecting the model's loss. These later layers exhibited abnormally high activation norms while contributing minimally to the model's predictive cross-entropy as seen in the logit lens analysis below. By removing these layers and keeping the unembedding layer, we create Evo 2 20B without additional training.
Logit lens analysis of Evo 2 40B. Orange: mean embedding L2 norm by block. Blue: excess cross-entropy (bits) relative to the final layer. The plateau in cross-entropy beyond block ~20 indicates that later layers contribute minimally to next-token prediction, while their activation norms spike dramatically.