-
Notifications
You must be signed in to change notification settings - Fork 0
add hw on pandas #12
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
base: main
Are you sure you want to change the base?
add hw on pandas #12
Changes from all commits
File filter
Filter by extension
Conversations
Jump to
Diff view
Diff view
There are no files selected for viewing
Large diffs are not rendered by default.
Large diffs are not rendered by default.
| Original file line number | Diff line number | Diff line change | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| @@ -0,0 +1,149 @@ | ||||||||||||||||||||||
| import pandas as pd | ||||||||||||||||||||||
| import numpy as np | ||||||||||||||||||||||
| import seaborn as sns | ||||||||||||||||||||||
| import matplotlib.pyplot as plt | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # read dataframe | ||||||||||||||||||||||
| df = pd.read_csv('../data/train.csv') | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # Drawing histograms | ||||||||||||||||||||||
| sns.set() | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| fig, ax = plt.subplots(2, 2, figsize=(12, 10)) | ||||||||||||||||||||||
| fig.suptitle('Distributions nucleotides over positions in reads', fontsize=18) | ||||||||||||||||||||||
| ax[0, 0].hist(df['A'].dropna(), bins=30) | ||||||||||||||||||||||
| ax[0, 1].hist(df['C'].dropna(), bins=30) | ||||||||||||||||||||||
| ax[1, 0].hist(df['T'].dropna(), bins=30) | ||||||||||||||||||||||
| ax[1, 1].hist(df['G'].dropna(), bins=30) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| ax[0, 0].set_title('A', fontsize=16) | ||||||||||||||||||||||
| ax[0, 1].set_title('C', fontsize=16) | ||||||||||||||||||||||
| ax[1, 0].set_title('T', fontsize=16) | ||||||||||||||||||||||
| ax[1, 1].set_title('G', fontsize=16) | ||||||||||||||||||||||
|
Comment on lines
+13
to
+23
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more. Было бы удобно отобразить это на одном графике. Плохой вариант - наложенные гистограммы с прозрачностью. Более хороший - barplot с группировкой или stacked barplot |
||||||||||||||||||||||
|
|
||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # Selecting special rows and cols | ||||||||||||||||||||||
| matches_mean = df.matches.mean() | ||||||||||||||||||||||
| df_subset = df.query('matches > @matches_mean') | ||||||||||||||||||||||
| df_subset = df_subset[['pos', 'reads_all', 'mismatches', 'deletions', 'insertions']] | ||||||||||||||||||||||
| df_subset.to_csv('../data/train_part.csv', index=False, sep=',') | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # Read dataframe with pokemons | ||||||||||||||||||||||
| pokemons_df = pd.read_csv('../data/Pokemon.csv') | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # Show general info about dataframe | ||||||||||||||||||||||
| pokemons_df.info() | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # Renaming columns | ||||||||||||||||||||||
| new_columns = {'#': 'Id', 'Type 1': 'Type_1', 'Type 2': 'Type_2', 'Sp. Atk': 'Sp_atk', 'Sp. Def': 'Sp_def'} | ||||||||||||||||||||||
| pokemons_df.rename(columns=new_columns, inplace=True) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # Checking for missing values | ||||||||||||||||||||||
| print('Не стоит беспокоиться об NA') if pokemons_df.count().min() == pokemons_df.shape[0] else print('Oops!') | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| plt.figure(figsize=(10, 8)) | ||||||||||||||||||||||
| plt.title('Heatmap of NA values in dataframe', fontsize=18) | ||||||||||||||||||||||
| sns.heatmap(pokemons_df.isna()) | ||||||||||||||||||||||
| plt.xticks(fontsize=14) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| print('\tПеременная\tПроцент NA') | ||||||||||||||||||||||
| print('---------------------------------------') | ||||||||||||||||||||||
| for col in pokemons_df.columns: | ||||||||||||||||||||||
| percentage = pokemons_df[col].isna().mean()*100 | ||||||||||||||||||||||
| if percentage == 0: | ||||||||||||||||||||||
| percentage = int(percentage) | ||||||||||||||||||||||
| else: | ||||||||||||||||||||||
| percentage = round(percentage, 2) | ||||||||||||||||||||||
| print('{:>14} {:>14} %'.format(col, percentage)) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # Plotting Type 1 and Type 2 classes | ||||||||||||||||||||||
| plt.figure(figsize=(15, 8)) | ||||||||||||||||||||||
| plt.subplot(1, 2, 1) | ||||||||||||||||||||||
| sns.countplot(pokemons_df['Type_1']) | ||||||||||||||||||||||
| plt.title('Type 1 classes', fontsize=18) | ||||||||||||||||||||||
| plt.xlabel('Type 1', fontsize=14) | ||||||||||||||||||||||
| plt.ylabel('Count', fontsize=14) | ||||||||||||||||||||||
| plt.xticks(rotation=90) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| plt.subplot(1, 2, 2) | ||||||||||||||||||||||
| sns.countplot(pokemons_df['Type_2']) | ||||||||||||||||||||||
| plt.title('Type 2 classes', fontsize=18) | ||||||||||||||||||||||
| plt.xlabel('Type 2', fontsize=14) | ||||||||||||||||||||||
| plt.ylabel('Count', fontsize=14) | ||||||||||||||||||||||
| plt.xticks(rotation=90) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # Plotting Generation and Legendary classes | ||||||||||||||||||||||
| plt.figure(figsize=(15, 8)) | ||||||||||||||||||||||
| plt.subplot(1, 2, 1) | ||||||||||||||||||||||
| sns.countplot(pokemons_df.Generation) | ||||||||||||||||||||||
| plt.xlabel('Generation', fontsize=14) | ||||||||||||||||||||||
| plt.ylabel('count', fontsize=14) | ||||||||||||||||||||||
| plt.xticks(fontsize=14) | ||||||||||||||||||||||
| plt.yticks(fontsize=14) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| plt.subplot(1, 2, 2) | ||||||||||||||||||||||
| sns.countplot(pokemons_df.Legendary) | ||||||||||||||||||||||
| plt.xlabel('Legendary', fontsize=14) | ||||||||||||||||||||||
| plt.ylabel('count', fontsize=14) | ||||||||||||||||||||||
| plt.xticks(fontsize=14) | ||||||||||||||||||||||
| plt.yticks(fontsize=14) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # Plotting pair plots for numeric features | ||||||||||||||||||||||
| numeric_cols = ['Total', 'HP', 'Attack', 'Defense', 'Sp_atk', 'Sp_def', 'Speed'] | ||||||||||||||||||||||
| sns.pairplot(pokemons_df[numeric_cols]) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # Plotting boxplots for numeric features | ||||||||||||||||||||||
| plt.figure(figsize=(18, 10)) | ||||||||||||||||||||||
| for i, col in enumerate(numeric_cols): | ||||||||||||||||||||||
| plt.subplot(2, 4, i+1) | ||||||||||||||||||||||
| sns.boxplot(pokemons_df[col]) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # Plotting Pearson correlation heatmap for numeric features | ||||||||||||||||||||||
| plt.figure(figsize=(10, 8)) | ||||||||||||||||||||||
| sns.heatmap(pokemons_df[numeric_cols].corr(), annot=True) | ||||||||||||||||||||||
| plt.title('Pearson correlation', fontsize=18) | ||||||||||||||||||||||
| plt.xticks(fontsize=14) | ||||||||||||||||||||||
| plt.yticks(fontsize=14) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # functions for reading gff and bed formats | ||||||||||||||||||||||
| def read_gff(path_to_file): | ||||||||||||||||||||||
| rrna_annotation_df = pd.read_csv(path_to_file, header=1, sep='\t') | ||||||||||||||||||||||
| rrna_annotation_df = pd.DataFrame(np.r_[np.array(rrna_annotation_df.columns)[np.newaxis, :], | ||||||||||||||||||||||
| rrna_annotation_df.values]) | ||||||||||||||||||||||
| rrna_annotation_df.columns = ['chromosome', 'source', 'type', 'start', 'end', | ||||||||||||||||||||||
| 'score', 'strand', 'phase', 'attributes'] | ||||||||||||||||||||||
| rrna_annotation_df[['start', 'end']] = rrna_annotation_df[['start', 'end']].astype(int) | ||||||||||||||||||||||
| rrna_annotation_df['score'] = rrna_annotation_df['score'].astype(float) | ||||||||||||||||||||||
|
Comment on lines
+113
to
+119
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more. Я не совсем понял к чему весь этот огород с индексами. Чтобы убрать комменты?
Suggested change
|
||||||||||||||||||||||
|
|
||||||||||||||||||||||
| return rrna_annotation_df | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
|
|
||||||||||||||||||||||
| def read_bed6(path_to_file): | ||||||||||||||||||||||
| alignment_bed_df = pd.read_csv(path_to_file, sep='\t', header=None) | ||||||||||||||||||||||
| alignment_bed_df.columns = ['chromosome', 'start', 'end', 'name', 'score', 'strand'] | ||||||||||||||||||||||
| return alignment_bed_df | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # creating new column with RNA type | ||||||||||||||||||||||
| rrna_annotation_df = read_gff('../data/rrna_annotation.gff') | ||||||||||||||||||||||
| rrna_annotation_df['rna_type'] = rrna_annotation_df.attributes.apply(lambda x: x.split('=')[1].split('_')[0]) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # collecting statistics on rna types in each chromosome | ||||||||||||||||||||||
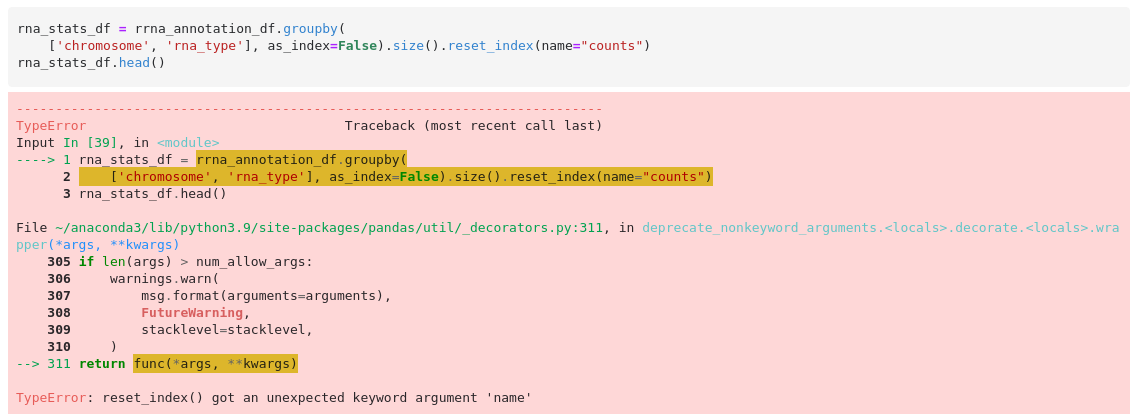
| rna_stats_df = rrna_annotation_df.groupby( | ||||||||||||||||||||||
| ['chromosome', 'rna_type'], as_index=False).size().reset_index(name='counts') | ||||||||||||||||||||||
|
Comment on lines
+135
to
+136
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more. |
||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # plotting barplot for different RNAs on each chromosome | ||||||||||||||||||||||
| plt.figure(figsize=(12, 10)) | ||||||||||||||||||||||
| sns.barplot(x='counts', y='chromosome', hue='rna_type', data=rna_stats_df) | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # reading bed file | ||||||||||||||||||||||
| alignment_bed_df = read_bed6('../data/alignment.bed') | ||||||||||||||||||||||
|
|
||||||||||||||||||||||
| # intersection (analogue of bedtools intersect command) | ||||||||||||||||||||||
| df_intersected = rrna_annotation_df.merge(alignment_bed_df, on=['chromosome'], how='inner') | ||||||||||||||||||||||
| is_inside = df_intersected[['start_x', 'end_x', 'start_y', 'end_y']].apply( | ||||||||||||||||||||||
| lambda x: x[0] > x[2] and x[1] < x[3], axis=1) | ||||||||||||||||||||||
|
Comment on lines
+147
to
+148
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more. Можно было просто сделать сравнения между колонками, но так тоже можно |
||||||||||||||||||||||
| df_intersected = df_intersected[is_inside] | ||||||||||||||||||||||
There was a problem hiding this comment.
Choose a reason for hiding this comment
The reason will be displayed to describe this comment to others. Learn more.
Пути к файлам лучше всегда указывать через
os.path.join