-
Notifications
You must be signed in to change notification settings - Fork 0
create hw with classes #11
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
base: main
Are you sure you want to change the base?
Changes from all commits
File filter
Filter by extension
Conversations
Jump to
Diff view
Diff view
There are no files selected for viewing
Large diffs are not rendered by default.
Large diffs are not rendered by default.
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,185 @@ | ||
| from Bio.Seq import Seq | ||
| from Bio import SeqIO | ||
| from Bio.SeqUtils import GC | ||
| import numpy as np | ||
| import seaborn as sns | ||
| import matplotlib.pyplot as plt | ||
|
|
||
|
|
||
| class Employee: | ||
|
|
||
| def __init__(self, name, age, work_experience): | ||
| self.name = name | ||
| self.age = age | ||
| self.work_experience = work_experience | ||
|
|
||
| def __str__(self): | ||
| return 'Information about employee: {}'.format(self.name) | ||
|
|
||
| def proposed_position(self): | ||
| ''' | ||
| Shows the position the candidate is applying for | ||
| :return: str, position name | ||
| ''' | ||
| if self.work_experience <= 1: | ||
| position = 'Junior DS' | ||
| elif 1 < self.work_experience <= 3: | ||
| position = 'Middle DS' | ||
| else: | ||
| position = 'Senior DS' | ||
| return position | ||
|
|
||
| def proposed_salary(self, position): | ||
| ''' | ||
| Shows salary the candidate wants to have | ||
| :param position: str, name of the position | ||
| :return: int, salary in dollars | ||
| ''' | ||
| if position == 'Junior DS': | ||
| salary = 1000 | ||
| elif position == 'Middle DS': | ||
| salary = 5000 | ||
| else: | ||
| salary = 10000 | ||
| return salary | ||
|
|
||
| def view_info(self): | ||
| ''' | ||
| Shows basic information about candidate | ||
| :return: str with information | ||
| ''' | ||
| position = self.proposed_position() | ||
| salary = self.proposed_salary(position) | ||
| return f"Name: {self.name}\nAge: {self.age} years\nPosition: {position}\nSalary {salary} $" | ||
|
|
||
|
|
||
| class SetModified(set): | ||
|
|
||
| def __init__(self, *args): | ||
| self.positive_numbers = [arg for arg in args if isinstance(arg, (int, float)) and arg > 0] | ||
| super().__init__(self.positive_numbers) | ||
|
|
||
| def add(self, element): | ||
| if isinstance(element, (int, float)) and element > 0: | ||
| super().add(element) | ||
|
|
||
| def update(self, *elements): | ||
| for element in elements: | ||
| self.add(element) | ||
|
|
||
|
|
||
| class RNA: | ||
|
|
||
| def __init__(self, rna): | ||
| self.rna = Seq(rna) | ||
|
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more. |
||
|
|
||
| def translation(self): | ||
| ''' | ||
| Translates the RNA sequence into protein | ||
| :return: protein sequence | ||
| ''' | ||
| return self.rna.translate() | ||
|
|
||
| def back_transcription(self): | ||
| ''' | ||
| Convert RNA sequence into DNA | ||
| :return: DNA sequence | ||
| ''' | ||
| return self.rna.back_transcribe() | ||
|
|
||
|
|
||
| class FastaStats: | ||
|
|
||
| def __init__(self, path): | ||
| self.path = path | ||
|
|
||
| def __str__(self): | ||
| return "Path to the file: {}".format(self.path) | ||
|
|
||
| def count_sequences(self): | ||
| ''' | ||
| Counts number of sequences presented in the file | ||
| :return: int, number of sequences | ||
| ''' | ||
| seq_numb = 0 | ||
| with open(self.path) as f: | ||
| for record in SeqIO.parse(f, "fasta"): | ||
|
Comment on lines
+105
to
+106
There was a problem hiding this comment. Choose a reason for hiding this commentThe reason will be displayed to describe this comment to others. Learn more. Ты считываешь данные в каждой функции. Наверное, лучше выделить это в отдельный метод и считать всё один раз в конструкторе |
||
| seq_numb += 1 | ||
| return seq_numb | ||
|
|
||
| def calc_sequences_length(self): | ||
| ''' | ||
| Calculates length of sequences presented in the file | ||
| :return: list with lengths of sequences | ||
| ''' | ||
| with open(self.path) as f: | ||
| length_ls = [len(record.seq) for record in SeqIO.parse(f, "fasta")] | ||
| return length_ls | ||
|
|
||
| def calc_gc_content_per_seq(self): | ||
| ''' | ||
| Calculates GC content of sequences presented in the file | ||
| :return: list with GC content per each sequence | ||
| ''' | ||
| with open(self.path) as f: | ||
| gc_content_ls = [GC(record.seq) for record in SeqIO.parse(f, "fasta")] | ||
| return gc_content_ls | ||
|
|
||
| def calc_mean_gc_content(self): | ||
| ''' | ||
| Calculates mean GC content over all sequences presented in the file | ||
| :return: int, mean GC content (%) | ||
| ''' | ||
| return np.mean(self.calc_gc_content_per_seq()) | ||
|
|
||
| def calc_k_mers(self, kmer_size): | ||
| ''' | ||
| Counts k-mer over all sequences in the file | ||
| :param kmer_size: int, size of k-mers | ||
| :return: dictionary, where keys - kmers, values - their counts | ||
| ''' | ||
| k_mer_counts = {} | ||
| with open(self.path) as f: | ||
| for record in SeqIO.parse(f, "fasta"): | ||
| for i in range(len(record.seq)-kmer_size+1): | ||
| kmer = str(record.seq[i:i+kmer_size]) | ||
| k_mer_counts[kmer] = k_mer_counts.get(kmer, 0) + 1 | ||
| k_mer_counts = dict(sorted(k_mer_counts.items(), key=lambda item: item[1], reverse=True)) | ||
| return k_mer_counts | ||
|
|
||
| def plot_length_distribution(self): | ||
| ''' | ||
| Plots length distribution for sequences presented in the file | ||
| ''' | ||
| length_ls = self.calc_sequences_length() | ||
| plt.figure(figsize=(10, 8)) | ||
| sns.distplot(length_ls, bins=30) | ||
| plt.title('Length distribution', fontsize=15) | ||
| plt.xlabel('length', fontsize=15) | ||
| plt.ylabel('frequency', fontsize=15) | ||
|
|
||
| def plot_gc_content_distribution(self): | ||
| ''' | ||
| Plots GC content distribution for sequences presented in the file | ||
| ''' | ||
| gc_content_ls = self.calc_gc_content_per_seq() | ||
| mean_gc_content = self.calc_mean_gc_content() | ||
| plt.figure(figsize=(10, 8)) | ||
| sns.distplot(gc_content_ls, bins=30) | ||
| plt.title('GC content distribution', fontsize=15) | ||
| plt.xlabel('GC content, %', fontsize=15) | ||
| plt.ylabel('frequency', fontsize=15) | ||
| plt.axvline(mean_gc_content, color='red', linestyle='--') | ||
|
|
||
| def plot_kmer_distribution(self, kmer_size): | ||
| ''' | ||
| Plots frequency of k-mers over all sequences | ||
| :param kmer_size: int, size of k-mers | ||
| ''' | ||
| k_mer_counts = self.calc_k_mers(kmer_size=kmer_size) | ||
| plt.figure(figsize=(12, 10)) | ||
| plt.bar(k_mer_counts.keys(), k_mer_counts.values(), width=1) | ||
| plt.title('{}-mers distribution'.format(kmer_size), fontsize=15) | ||
| plt.xlabel('{}-mers'.format(kmer_size), fontsize=15) | ||
| plt.ylabel('counts', fontsize=15) | ||
| plt.xticks(fontsize=3, rotation=90) | ||
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,4 @@ | ||
| biopython==1.79 | ||
| matplotlib==3.1.3 | ||
| seaborn==0.10.0 | ||
| numpy==1.20.0 |
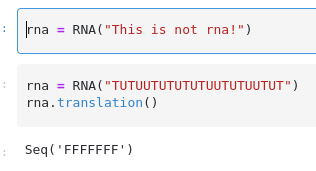
There was a problem hiding this comment.
Choose a reason for hiding this comment
The reason will be displayed to describe this comment to others. Learn more.
Записывать это в атрибут не нужно